POPULATION GENOMICS AT THE MCDONNELL GENOME INSTITUTE
I joined the Warren Lab at The Genome Institute (TGI) in June 2012 to undertake various research projects in population genomics. The Warren Lab specializes in de novo assembly methods while also assessing: 1) the evolution of paralogs in vertebrate and non-vertebrate genomes; 2) the influence of genetic variation on phenotypes as well as on transcriptional and protein networks; 3) the genetic roles in gene splicing and structural variation; and 4) the population genomics of genic regions and regulatory regions.
I've analyzed genomic data from a number of taxa, including cavefish, tsetse fly, Leishmania, sandfly, killifish, bald eagle, Amazon molly, vervet monkey, and colugo. Apart from a small number of analyses on parasite genomics, I've primarily implemented projects that are less disease-centric and would be categorized within the realm of population genetics and evolutionary genome science.
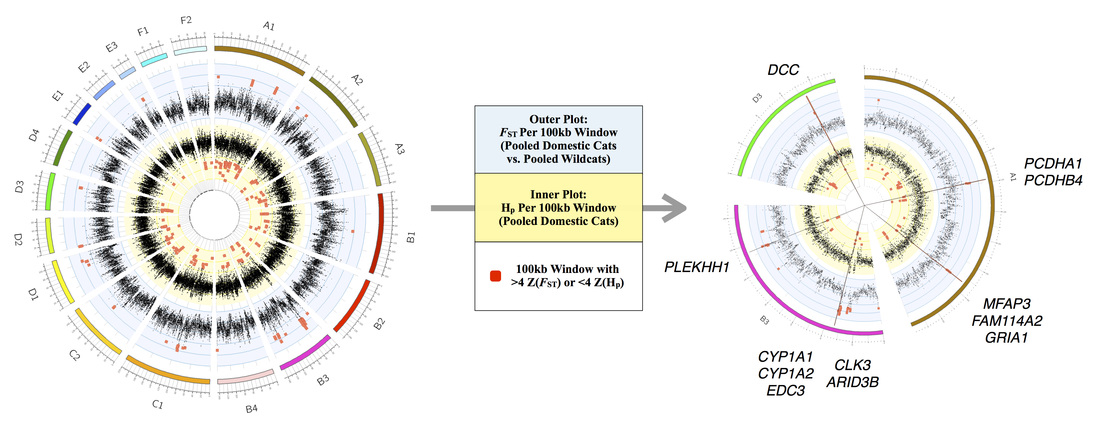
As a post-doctoral research associate, my main role at TGI involved leading a project to investigate the signatures of selection in the genomes of domestic cat breeds and their wildcat progenitor populations. This work, funded by grants from the National Human Genome Research Institute (NHGRI), was conducted in collaboration with Bill Murphy (Texas A&M) and Leslie Lyons (University of Missouri) and led to a recent publication in the Proceedings of the National Academy of Science.
I joined the Warren Lab at The Genome Institute (TGI) in June 2012 to undertake various research projects in population genomics. The Warren Lab specializes in de novo assembly methods while also assessing: 1) the evolution of paralogs in vertebrate and non-vertebrate genomes; 2) the influence of genetic variation on phenotypes as well as on transcriptional and protein networks; 3) the genetic roles in gene splicing and structural variation; and 4) the population genomics of genic regions and regulatory regions.
I've analyzed genomic data from a number of taxa, including cavefish, tsetse fly, Leishmania, sandfly, killifish, bald eagle, Amazon molly, vervet monkey, and colugo. Apart from a small number of analyses on parasite genomics, I've primarily implemented projects that are less disease-centric and would be categorized within the realm of population genetics and evolutionary genome science.
As a post-doctoral research associate, my main role at TGI involved leading a project to investigate the signatures of selection in the genomes of domestic cat breeds and their wildcat progenitor populations. This work, funded by grants from the National Human Genome Research Institute (NHGRI), was conducted in collaboration with Bill Murphy (Texas A&M) and Leslie Lyons (University of Missouri) and led to a recent publication in the Proceedings of the National Academy of Science.
Research into primate population genomics served as a secondary focus, where my work with the genomes of cynomologous macaque, capuchin, vervet monkey, chimpanzee, bonobo, and tarsier allowed me to re-tool pipelines and develop new analytic methods to most effectively and efficiently make use of large-scale genomics data. I have recently implemented procedures to 1) assess positive selection and 2) detect gene family expansions and contractions. I'm currently working with developers of tools for rapid and scalable detection of selective sweeps in datasets using whole genome sequence generated from population samples.
While much of this work will become published in the near future, I also spend some of my time conducting formal and informal mentoring in biology, population genetics, genomics, and bioinformatics for undergraduate, post-baccelaureate and graduate students.